
qScript Ultra SuperMix
- Enhanced Thermostability – Engineered for RT activity up to 65°C, overcoming challenging secondary structures
- Superior Speed – 10 minute reaction time with a total protocol time of 13 minutes
- Maximum Yield & Sensitivity – Wide dynamic range with no loss in cDNA abundance linearity from total RNA 2.5 µg – 1 pg
- Ultimate Inhibitor Resistance – Overcome a wide array of PCR inhibitors (salt, heparin, melanin, etc.)
- Balanced Coverage of Long Transcripts – Unbiased representation of the transcriptome in cDNA product
- Ease of Use – Single tube mix with all required components for cDNA synthesis except RNA template
qScript Ultra SuperMix is intended for molecular biology applications. This product is not intended for the diagnosis, prevention or treatment of a disease.
qScript Ultra SuperMix
Description
qScript Ultra SuperMix is a highly stabilized, efficient and easy-to-use single tube master mix for the synthesis of first-strand cDNA to reverse transcribe RNA to cDNA. A key component is a novel, state-of-the-art, RNase H deficient reverse transcriptase that was engineered for improved thermostability, velocity, processivity, and resistance to many common inhibitors. qScript Ultra SuperMix contains all required components for first-strand cDNA synthesis except RNA template and is directly compatible with downstream 2-step RT-qPCR or RT-PCR procedures.
Details
|
|
95217-025 |
95217-100 |
95217-500 |
| 5X qScript Ultra SuperMix |
1 x 100 µl |
1 x 400 µl |
1 x 2 mL |
|
Optimized master mix containing buffer, magnesium, dNTPs, optimized primer mix and qScript Ultra RT |
|||
Performance Data
Comparison of protocol length for first-strand cDNA synthesis kits.
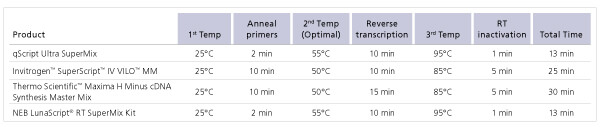
Comparison of protocol length for first-strand cDNA synthesis kits. Incorporating the highly rapid and processive qScript Ultra reverse transcriptase,
the qScript Ultra SuperMix requires 10 minutes for first-strand cDNA synthesis over a broad range of input RNA quantities.
High Range Input Linearity.
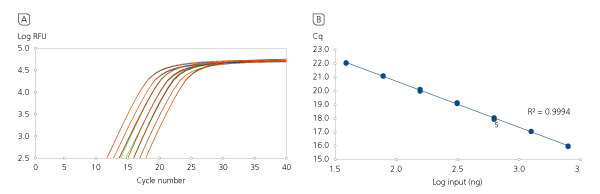
High Range Input Linearity. First-strand cDNA synthesis reactions were carried out using increasing quantities of total human RNA input, followed by analysis of a portion of the products by beta-actin qPCR. In only a 10 minute reaction time, the qScript Ultra SuperMix accommodates up to 2.5 μg total RNA input without a loss in cDNA abundance linearity.
Low input reaction sensitivity.
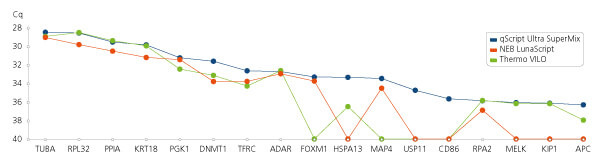
Low input reaction sensitivity. First-strand cDNA synthesis reactions were assembled and reacted with 2 pg total human RNA, then a portion (15%) of the reaction was used for qPCR analysis. The Cq values of duplicate reactions were averaged and plotted. Where no amplification took place, values were plotted as Cq = 40.
Viral RNA cDNA synthesis in the presence of carrier RNA.
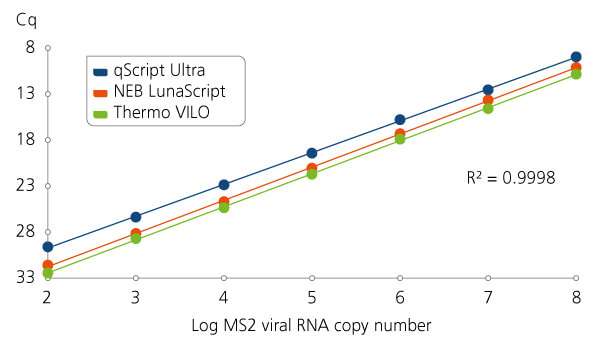
Viral RNA cDNA synthesis in the presence of carrier RNA. First-strand cDNA synthesis reactions were assembled and reacted with the indicated quantities of MS2 viral RNA in the presence of 1 μg poly(A), which is a common carrier that can co-purify with viral extraction preparations and potentially react and interfere with detection. qScript Ultra SuperMix displays higher yields of cDNA after qPCR detection of a portion of the viral material.
Inhibitor resistance across wide range of sample types.
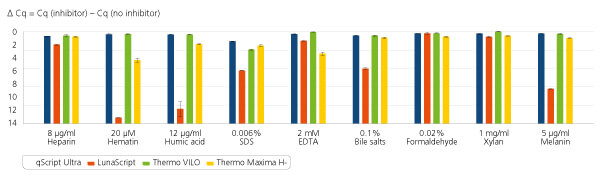
Inhibitor resistance across wide range of sample types. Standard, randomer-primed first-stand cDNA synthesis reactions were carried out using 50 ng total human RNA in the presence of several common reaction inhibitors. Following cDNA synthesis, both products and inhibitors were diluted such that 1/50 of the initial reaction was used for qPCR. Cq values were compared with reactions without inhibitors to show relative inhibitor tolerance in the first-strand reaction.
Balanced coverage of long transcripts
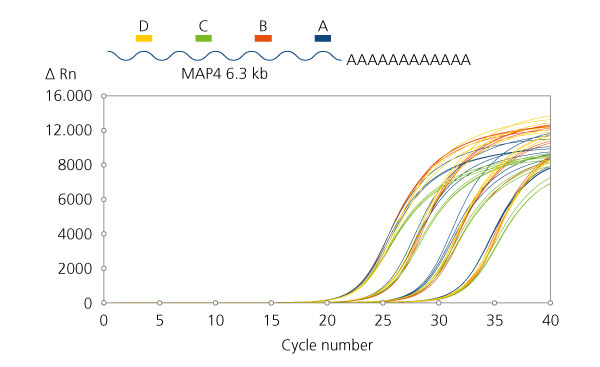
Balanced coverage of long transcripts. qPCR primer assays were designed at several intervals along the length of the 6.3 kb MAP4 mRNA transcript. First strand cDNA synthesis using qScript Ultra SuperMix was performed with total RNA input levels of 100 ng, 10 ng, 1 ng and 100 pg. Equivalent Cq calls for 4 different regions of the 6.3 kb transcript spanning the 3′-end near the poly A tail to the 5′-end transcriptional start side, at each input amount of UHR total RNA. The “A” assay refers to the red curves, “B” assay refers to the blue curves, “C” assay refers to the green curves, and “D” assay refers to the yellow curves.
Documents & Downloads
Customer Product Reviews
| 5 star | 77% | |
| 4 star | 20% | |
| 3 star | 0% | |
| 2 star | 0% | |
| 1 star | 1% |
 qScript Ultra SuperMix
qScript Ultra SuperMix

This is a great alternative to SuperScript IV VILO, highly recommend using this!
Super easy to use, quick and simple.
qScript was comparable to the BioRad product iScript