Extracta DNA Prep for PCR
- Simple, reagent-based system requires minimal technical skill
- Incubation step can be carried out in 96-well PCR plates or tubes using a standard DNA thermal cycler
- Compatible with a wide-range of clinical specimens, plant and animal tissues, and environmental samples
- Optional stabilization buffer allows for extended storage of extracted DNA templates
Extracta DNA Prep for PCR is intended for molecular biology applications. This product is not intended for the diagnosis, prevention or treatment of a disease.
Extracta DNA Prep for PCR
Description
Extracta DNA Prep for PCR is a two-component reagent kit for rapid extraction of PCR-ready genomic DNA from a variety of tissues. Samples are processed in less than 30 minutes with minimal hands-on time and technical skill. Extracted genomic DNA is suitable for sensitive downstream PCR applications including end-point PCR, High Resolution Melt Analysis (HRM) and quantitative real-time PCR (qPCR) without requiring any additional clean-up. In addition, the extracted DNA may be
used in multiplexed PCR applications such as transgene or knock-out analyses. Tissue extractions can be done in tubes, plates or deep-well blocks to allow for adaptation to workflow and automation on liquid-handling workstations.
Details
- Extraction Reagent (1 x 25 mL or 2 x 125 mL)
- Stabilization Buffer (1 x 25 mL or 2 x 125 mL)
Performance Data
Multiplex Genotyping
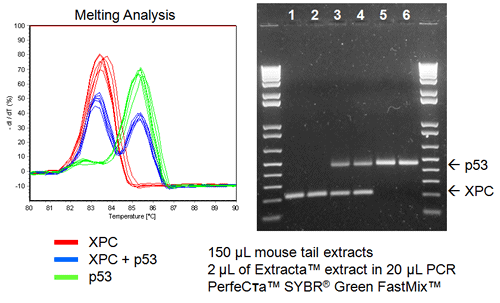
Multiplex Genotyping
Protocol Speed
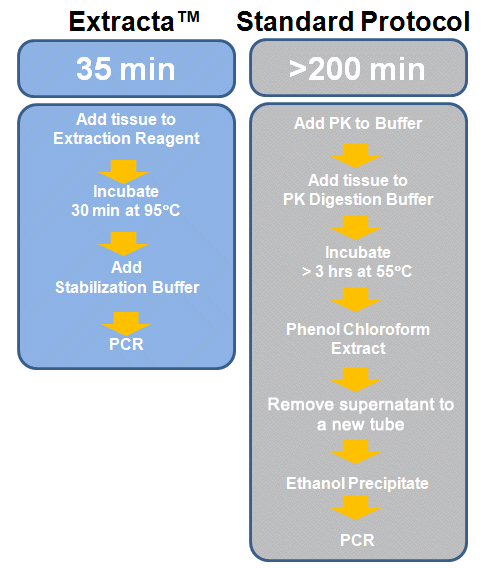
Extracta reduces time to results with a rapid, convenient protocol.
Comparison of Extract Yield: Mouse Ears
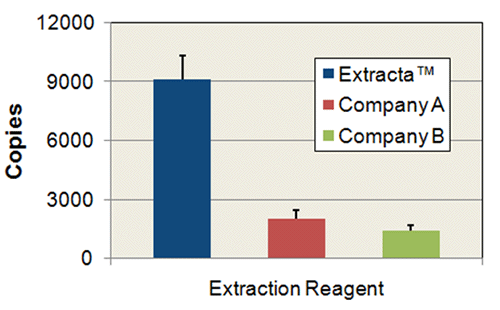
1 mm diameter mouse ear punches were extracted in 100 µL of Extracta DNA Prep for PCR – Tissue or reagents from competitor products according to each manufacturers instructions. 5.0 µL of each extract was used in 20 µL PCR reactions of mouse brain-derived neurotropic factor using PerfeCTa SYBR Green FastMix. PCR data was obtained in triplicate and converted to number of copies of target detected using a standard curve generated from purified mouse genomic DNA.
Comparison of Extract Yield: Mouse Tails
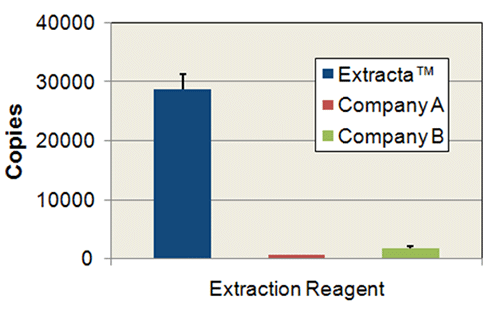
2 mm tail snips were extracted in 150 µL of Extracta DNA Prep for PCR – Tissue or reagents from competitor products according to each manufacturers instructions. 1.5 µL of each extract was used in 20 µL PCR reactions of mouse brain-derived neurotropic factor using PerfeCTa SYBR Green FastMix. PCR data was obtained in triplicate and converted to number of copies of target detected using a standard curve generated from purified mouse genomic DNA.
Documents & Downloads
Customer Product Reviews
| 5 star | 77% | |
| 4 star | 22% | |
| 3 star | 0% | |
| 2 star | 0% | |
| 1 star | 0% |
 Extracta DNA Prep for PCR
Extracta DNA Prep for PCR

It’s nice to have the reagents split up into smaller aliquots, it’s also helpful not to have to thaw the reagents because they can be stored at room temperature, unlike the reagents that we previously used.