
repliQa HiFi ToughMix
- High Fidelity – >90x wild type Taq
- Extreme Speed – up to 3x faster PCR results with extension rates as fast as 1 kb/sec*
- Tough Tested – tolerant to a wide range of PCR inhibitors
- Superior Sensitivity – higher yields from lower inputs as little as 2 pg
- Long Range – amplify +24 kb gDNA, +40 kb λDNA
*for fragments less than 1 kb in size
repliQa HiFi ToughMix is intended for molecular biology applications. This product is not intended for the diagnosis, prevention or treatment of a disease.
repliQa HiFi ToughMix
Description
The repliQa HiFi ToughMix is a 2x, ready-to-use master mix that contains all the components for high fidelity PCR amplification, including a genetically modified DNA polymerase coupled with hot start antibodies. This unique, next generation master mix provides 90x higher fidelity compared to Taq, while reducing time to PCR results by 2-3x. The extreme speed is enabled by extension times as fast as 1-10 kb/sec depending on target length. The enzyme is coupled with the industry leading ToughMix which is tolerant to a wide variety of inhibitors making it suitable for routine PCR, cloning, amplicon sequencing and site directed mutagenesis.
Details
2x reaction buffer containing optimized concentrations of MgCl2, dNTP’s and proprietarily formulated HiFi polymerase, hot start antibodies and ToughMix chemistry
Performance Data
Extreme Speed: Up to 3x faster results
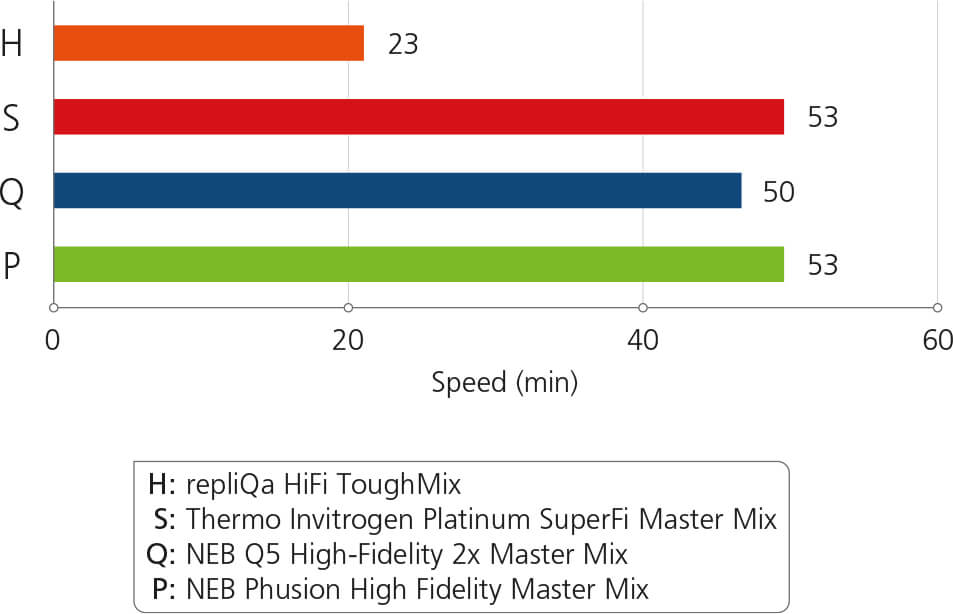
Comparison of speed. A 2 kb fragment was amplified in 50 μl reaction volumes according to the recommended protocol. Following a 30 s activation at 98°C; 30 cycles of PCR were performed: 98°C, 10 s; 60°C, 10 s; 68°C, 5–30 s. The thermal cycler had a ramp rate of 5°C/s.
Tough Tested: Tolerant to a wide range of PCR inhibitors
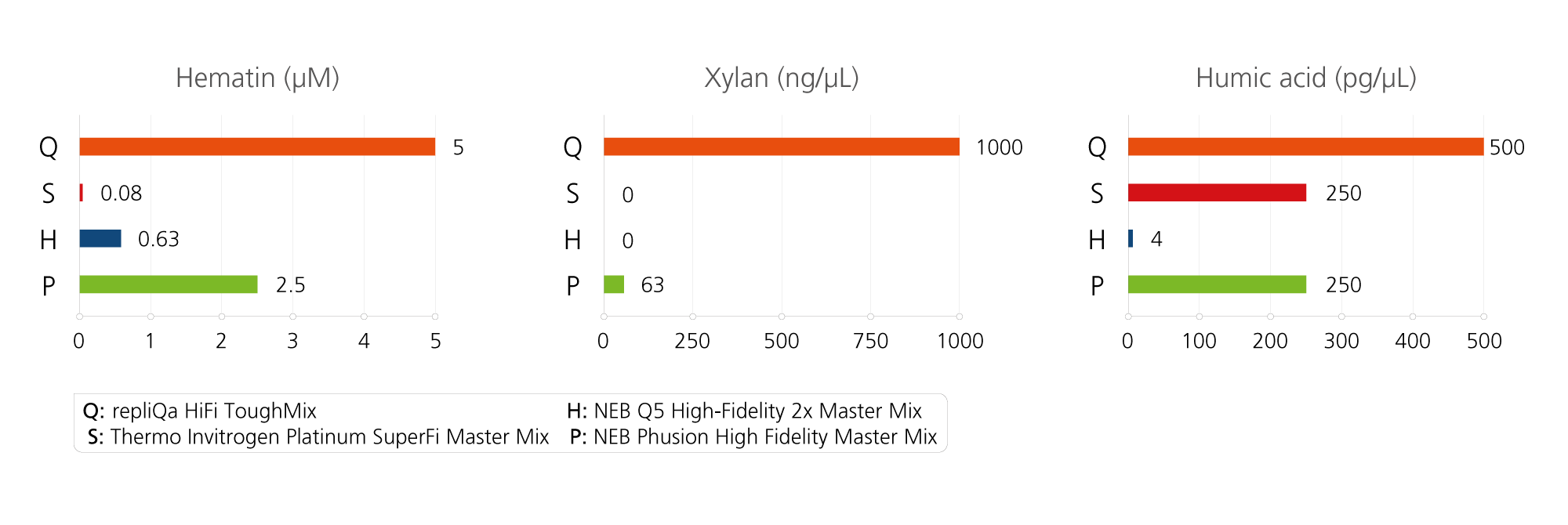
Strong Inhibitor Resistance. A 2 kb λ DNA template was amplified using each manufacturer’s recommended cycling conditions with different amounts io inhibitors. This experiment was run in duplicate.
Superior Yield and Sensitivity
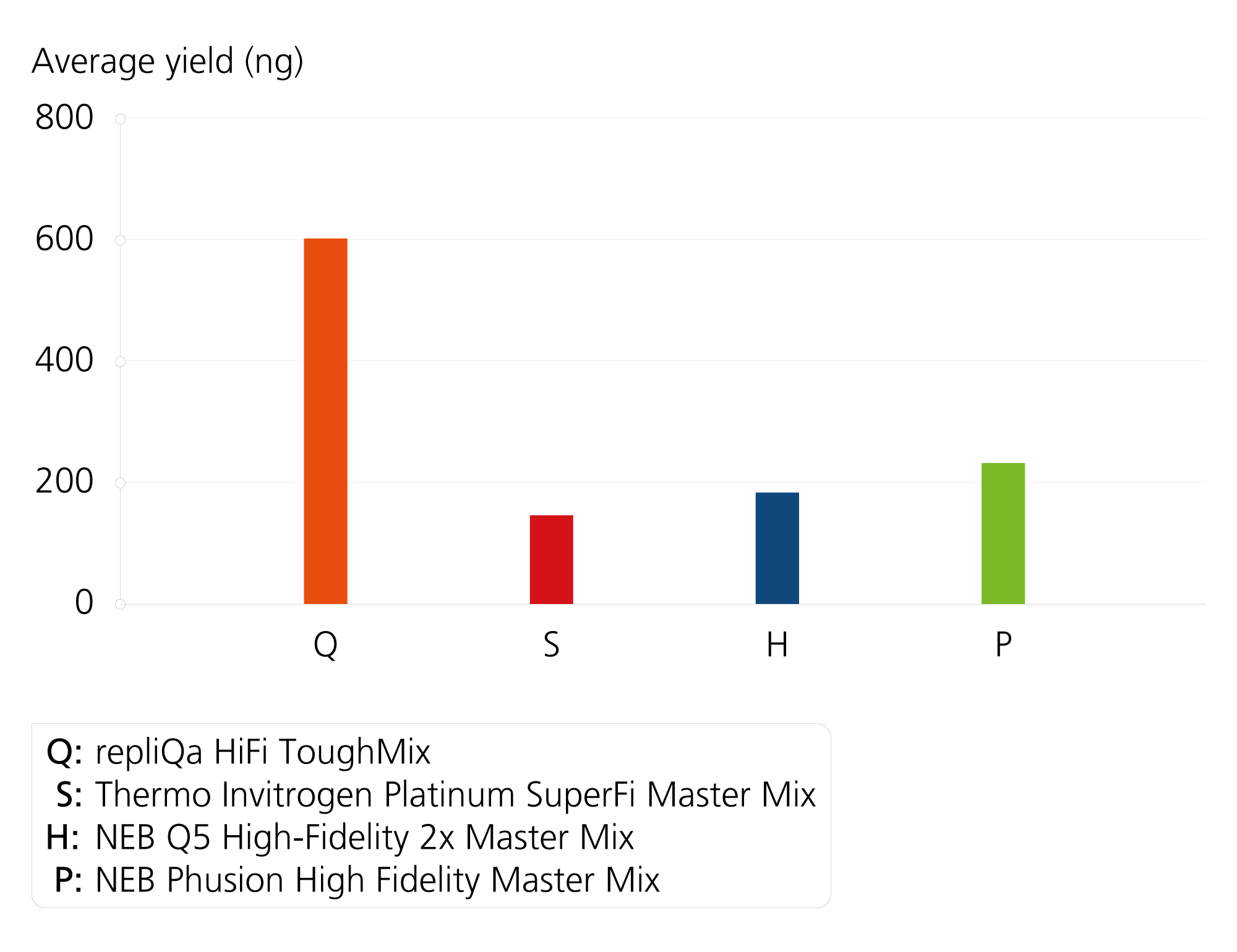
Comparison of yield. A gDNA template was amplified with varying GC-content and length targets using each manufacturer’s recommended cycling conditions. The experiment was run using 8 different targets, in duplicate.
Long Amplification and Consistent GC Tolerance
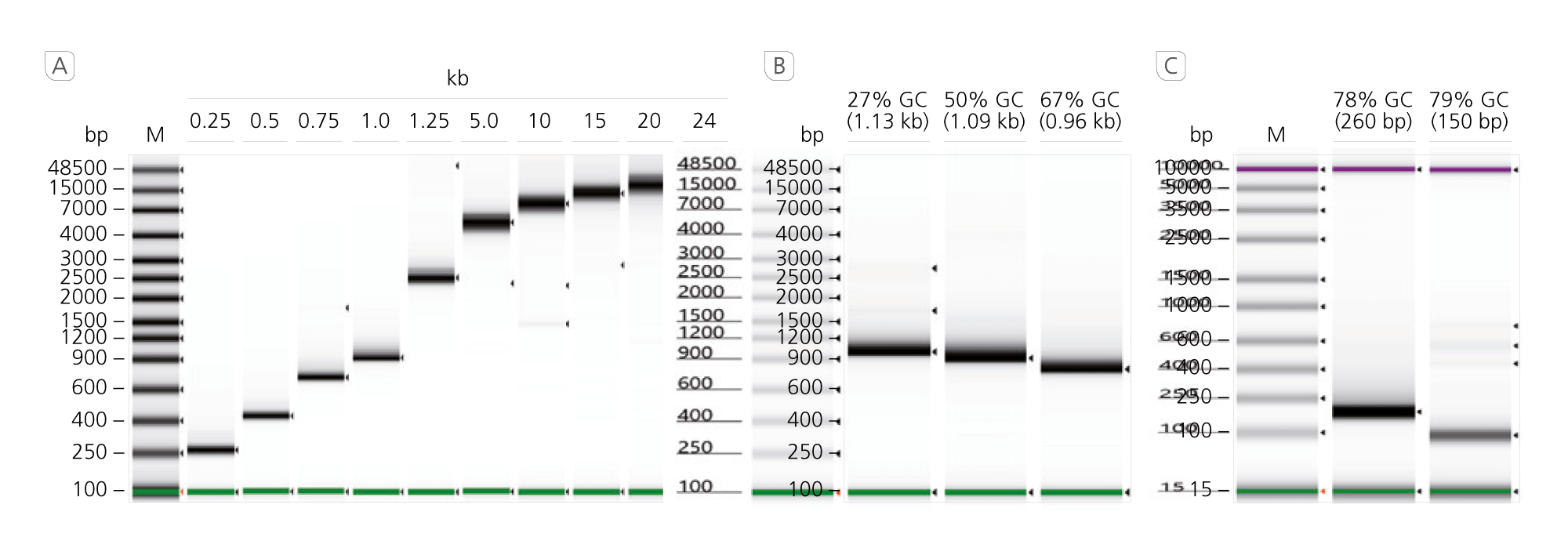

A Long Range Capabilities. A range from 0.25 to 24 kb of gDNA templates were amplified with varying GC-content and lengths using recommended cycling
conditions. B/C Wide GC-content tolerance range. gDNA templates were amplified with varying GC-content and lengths using recommended cycling conditions.
Documents & Downloads
Customer Product Reviews
| 5 star | 65% | |
| 4 star | 28% | |
| 3 star | 4% | |
| 2 star | 1% | |
| 1 star | 0% |
 repliQa HiFi ToughMix
repliQa HiFi ToughMix
Customer Testimonials
Welcome to the Quantabio webshop!
Please complete the new user account registration form
Once your account is setup, you'll be able to purchase any of the products on our site at any time with next day delivery for in stock products.


Worked on our dirty fecal samples.